This guide builds a Retrieval Augmented Generation pipeline that runs entirely on your hardware. No OpenAI key, no Anthropic key, no telemetry. PostgreSQL with pgvector stores the embeddings and the retrieval index. Ollama runs both the embedding model and the chat model. Around 250 lines of Python wires it together.
Most “self-hosted RAG” tutorials quietly call OpenAI for embeddings. That ships every chunk of every document you ingest to a third party and locks you into pricing and model choices you do not control. The pipeline here uses nomic-embed-text (768 dimensions, 274 MB) for embeddings and llama3.1:8b (Q4_K_M, 4.9 GB) for generation. Both run under Ollama. The vector store is plain Postgres 17 with the pgvector extension and a tsvector full text column for hybrid retrieval.
What you will end up with: a small Python program that ingests a folder of Markdown or text files, embeds and stores them, and answers questions with citations to the source documents. We test it on 50 articles from this site as the corpus.
Tested on: Ubuntu 22.04 LTS · PostgreSQL 17.9 · pgvector 0.8.0 · Ollama 0.23.1 · Python 3.10 · NVIDIA RTX 4090 (CPU inference also covered) · Verified 2026-05.
Why self host the whole RAG stack
RAG with a hosted embedding API is the default in most tutorials because it is the path of least friction. The hidden costs add up quickly once you move past the toy demo:
- Privacy. Every chunk of every document goes to a third party for embedding. Customer support tickets, internal wikis, code review history, contracts. Once it leaves your network, you have lost control of what happens next.
- Cost at scale. A modest 50,000 document corpus at 800 tokens per chunk runs about 40 million tokens. OpenAI text-embedding-3-small at $0.02 per 1M tokens is $0.80 to ingest once. Re-embedding when you switch chunk size, or running ingestion in dev and prod, multiplies that. Then every query embeds the question too.
- Lock in. The hosted embedding model gets deprecated on the provider’s schedule. You re-embed everything to migrate. Self hosting means you choose when to upgrade, and you can run two models side by side during the transition.
- Latency. A round trip to the API for every retrieval adds 100 to 300 ms before any DB work. Local Ollama on the same host returns embeddings in 30 to 60 ms.
The pipeline below has no API keys anywhere. You can airgap the host and it keeps working.
Architecture
RAG is four stages in a row.
Documents (md/pdf/html)
│
│ chunk + embed via Ollama
▼
PostgreSQL + pgvector
├─ chunks(content, embedding vector(768))
├─ HNSW index on embedding (cosine)
└─ GIN index on tsvector(content) for full text
│
│ hybrid retrieve: cosine top-K + tsvector top-K, fuse with RRF
▼
Llama 3.1 8B via Ollama
│ prompt = system rules + numbered context + question
▼
Answer with citations [1] [2] [3]The choices are deliberate:
- pgvector, not Chroma or FAISS. Postgres is already the operational database in many environments. pgvector lets you join embeddings to your application data in one query. Backups, replication, point in time recovery, and pg_stat_statements all work the same way they always have. A separate vector DB is a separate operational story to learn and own.
- Ollama for both embed and chat. One service to run, one set of models to keep up to date, one auth surface, one observability target. The Python SDK is the same for both calls.
- nomic-embed-text plus Llama 3.1 8B. The smallest pair that holds up on a real technical corpus. Both are open source. nomic-embed-text produces 768 dimensions and runs at hundreds of texts per second on a small GPU. Llama 3.1 8B at Q4 quant fits in 6 GB of VRAM. We discuss when to graduate to bigger models near the end.
- Hybrid retrieval, not pure vector. Pure cosine similarity has a known weakness on technical corpora: it matches semantic patterns better than exact tokens like product names, flag names, or version numbers. Section 6 shows a real failure on this corpus and section 7 fixes it.
Prerequisites
Three components that this guide does not install from scratch, with links to the dedicated guides:
- PostgreSQL 17. Follow Install PostgreSQL 17 on Rocky Linux 10 or on Ubuntu 24.04.
- pgvector 0.8.0 or newer. The HNSW index and the operators we rely on are in 0.5.0 and later, but 0.8.0 ships meaningful build performance improvements. See Install pgvector on PostgreSQL.
- Ollama 0.23.1 or newer. Earlier versions had a different embed API surface. Install Ollama on Rocky Linux or Ubuntu.
Pull the two models you need:
ollama pull nomic-embed-text
ollama pull llama3.1:8bVerify the embed API is reachable and returns the expected dimension:
$ curl -s http://127.0.0.1:11434/api/embed \
-d '{"model":"nomic-embed-text","input":"hello"}' | jq '.embeddings[0] | length'
768Create a database and role for the RAG app, and enable the vector extension:
sudo -u postgres psql <<'SQL'
CREATE ROLE rag LOGIN;
CREATE DATABASE rag OWNER rag;
\c rag
CREATE EXTENSION vector;
SQLSet up a Python virtual environment and install the four dependencies we need:
python3 -m venv .venv
. .venv/bin/activate
pip install "psycopg[binary]>=3.2" "pgvector>=0.3.6" "ollama>=0.4" tqdmThat is the whole stack. No LangChain, no LlamaIndex, no separate vector database, no API keys.
Database schema and indexes
Two tables. documents tracks one row per source file with a content hash for idempotent re ingestion. chunks holds one row per text fragment with its embedding and a generated tsvector column for full text search.
CREATE TABLE IF NOT EXISTS documents (
id BIGSERIAL PRIMARY KEY,
source TEXT NOT NULL UNIQUE,
title TEXT,
sha256 TEXT NOT NULL,
bytes INTEGER NOT NULL,
ingested_at TIMESTAMPTZ DEFAULT NOW()
);
CREATE TABLE IF NOT EXISTS chunks (
id BIGSERIAL PRIMARY KEY,
document_id BIGINT NOT NULL REFERENCES documents(id) ON DELETE CASCADE,
chunk_idx INTEGER NOT NULL,
content TEXT NOT NULL,
tokens INTEGER NOT NULL,
embedding vector(768) NOT NULL,
content_tsv tsvector GENERATED ALWAYS AS (to_tsvector('english', content)) STORED,
UNIQUE (document_id, chunk_idx)
);
CREATE INDEX IF NOT EXISTS chunks_embedding_hnsw
ON chunks USING hnsw (embedding vector_cosine_ops)
WITH (m = 16, ef_construction = 64);
CREATE INDEX IF NOT EXISTS chunks_content_tsv_gin
ON chunks USING gin (content_tsv);A few choices worth calling out:
- Generated tsvector column, not a tsvector index over content directly. Postgres maintains
content_tsvautomatically on every insert and update. The GIN index sees a stable column and stays compact. The application never has to remember to keep them in sync. - HNSW with
m=16andef_construction=64. These are sane defaults from the pgvector docs.mis the maximum number of bidirectional links per node in the HNSW graph. Highermimproves recall but enlarges the index roughly linearly.ef_constructioncontrols how many candidates are considered when inserting each vector. Higher gives better quality at the cost of build time. - UNIQUE on
(document_id, chunk_idx). Idempotent re ingestion. If a source file changes, the parentdocumentsrow is deleted and re inserted. The CASCADE drops the old chunks. Re ingestion of an unchanged file is a no op (we compare sha256 first). - Vector dimension 768. Matches nomic-embed-text. If you switch to a different embedding model, change this and re ingest. There is no graceful migration. Embeddings from different models are not interchangeable.
You can confirm the schema landed correctly with \d chunks:
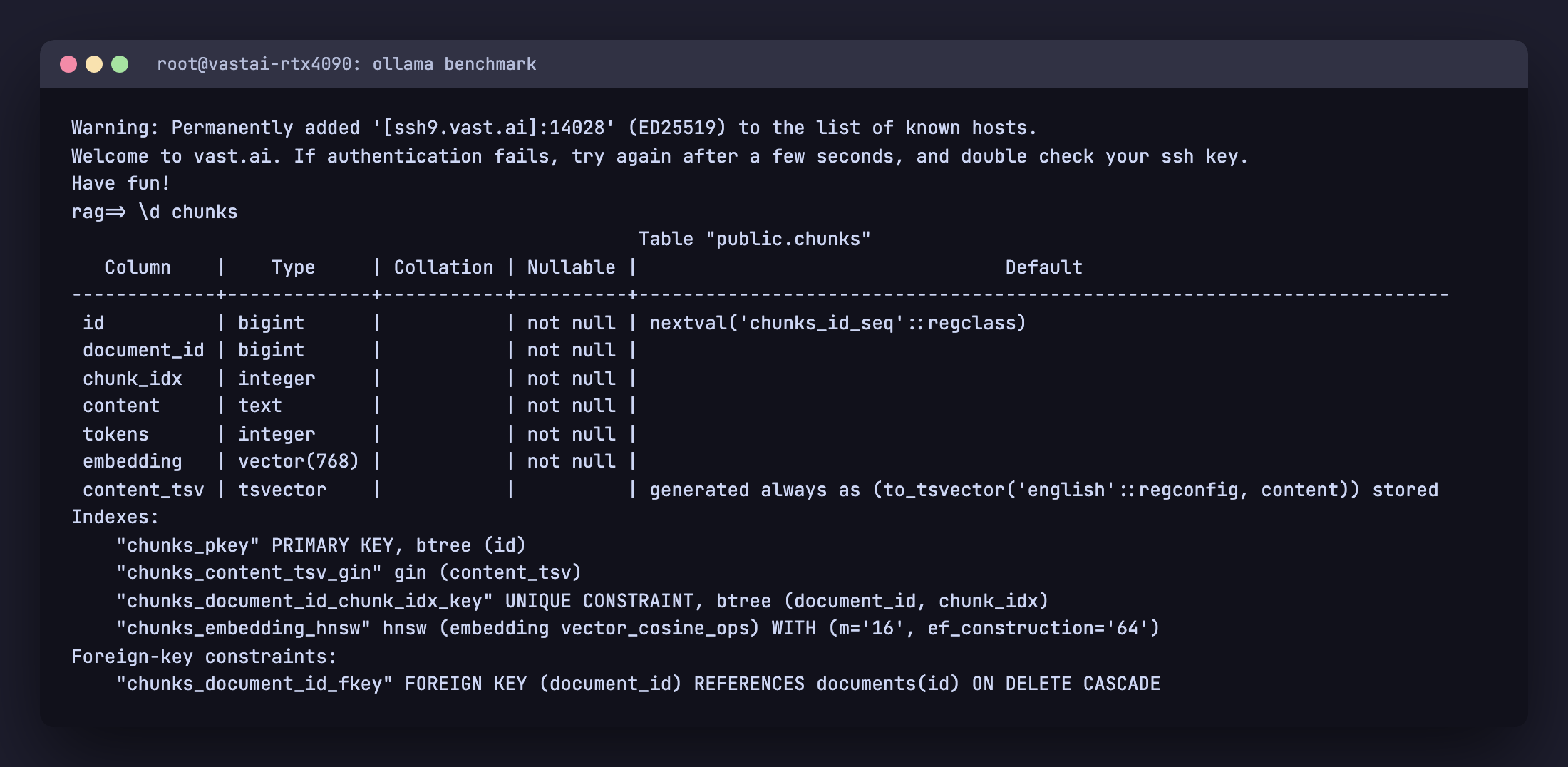
Ingestion pipeline
Ingestion does three things: chunk a document, embed each chunk, and insert the chunks. The whole pipeline lives in rag.py. The chunking function is the only piece that requires care.
Chunking
The aim is to split a document into pieces that are small enough that retrieval can be precise but large enough that each piece carries enough context to be useful in an answer. Around 800 tokens with 100 token overlap is a good default. We split paragraphs first, then sentences if a single paragraph is too big, then characters as a last resort. The overlap copies the tail of each chunk into the head of the next so an answer that straddles a chunk boundary still has continuous context.
TOKEN_RE = re.compile(r"\w+|[^\w\s]")
def tokens(text):
return len(TOKEN_RE.findall(text))
def split_text(text, target=800, overlap=100):
paras = re.split(r"\n\s*\n", text)
out, buf, buf_tokens = [], [], 0
for p in paras:
pt = tokens(p)
if buf_tokens + pt <= target:
buf.append(p); buf_tokens += pt
continue
if buf:
out.append("\n\n".join(buf))
if pt > target:
sents = re.split(r"(?<=[.!?])\s+", p)
sb, st = [], 0
for s in sents:
sti = tokens(s)
if st + sti > target and sb:
out.append(" ".join(sb)); sb, st = [], 0
sb.append(s); st += sti
if sb: out.append(" ".join(sb))
buf, buf_tokens = [], 0
else:
buf, buf_tokens = [p], pt
if buf:
out.append("\n\n".join(buf))
if overlap and len(out) > 1:
bridged = [out[0]]
for prev, cur in zip(out, out[1:]):
tail = " ".join(TOKEN_RE.findall(prev)[-overlap:])
bridged.append(tail + " " + cur)
return bridged
return outThe token counter uses a simple regex, not a real tokenizer. For chunking purposes the count just needs to be approximately right. Off by 5 percent is fine. Off by a factor of two is a bug.
Embedding and inserting
Ollama exposes an /api/embed endpoint that the Python SDK wraps. We embed one chunk per call here for clarity. In production, batch up to 32 or 64 chunks per call and parallelize across a few workers (see the production hardening section).
oclient = ollama.Client(host=os.environ.get("OLLAMA_HOST", "http://127.0.0.1:11434"))
def embed(texts):
out = []
for t in texts:
r = oclient.embed(model="nomic-embed-text", input=t)
out.append(r["embeddings"][0])
return outThe ingest function reads each file, computes a sha256, skips it if the database already has a row with the same source and the same hash, otherwise re inserts:
def cmd_ingest(directory):
files = [p for p in Path(directory).rglob("*.md") if not p.name.startswith(".")]
files += [p for p in Path(directory).rglob("*.txt") if not p.name.startswith(".")]
with db() as c, c.cursor() as cur:
for fp in sorted(files):
data = fp.read_bytes()
sha = hashlib.sha256(data).hexdigest()
source = str(fp.resolve())
cur.execute("SELECT id, sha256 FROM documents WHERE source = %s", (source,))
row = cur.fetchone()
if row and row[1] == sha:
continue # unchanged, skip
if row:
cur.execute("DELETE FROM documents WHERE id = %s", (row[0],))
cur.execute(
"INSERT INTO documents (source, title, sha256, bytes) "
"VALUES (%s, %s, %s, %s) RETURNING id",
(source, fp.stem, sha, len(data)),
)
doc_id = cur.fetchone()[0]
text = data.decode("utf-8", "replace").replace("\x00", "")
chs = split_text(text)
embs = embed(chs)
cur.executemany(
"INSERT INTO chunks (document_id, chunk_idx, content, tokens, embedding) "
"VALUES (%s, %s, %s, %s, %s)",
[(doc_id, i, ck, tokens(ck), e) for i, (ck, e) in enumerate(zip(chs, embs))],
)
c.commit()The register_vector call from the pgvector Python package teaches psycopg how to send Python lists as the vector type. Without it you get type errors on the executemany call.
Run it on a corpus. We use 50 articles from this site as a test set:
$ python rag.py ingest articles/
scanning articles/: 50 files
[5/50] ansible-lamp-lemp-stack.md (5 chunks)
[10/50] argocd-ingress-metallb.md (7 chunks)
...
[50/50] weave-gitops-install-migration-flux.md (5 chunks)
ingest done: 311 chunks, ~246147 tokens in 320.5s (1.0 chunks/s)320 seconds to embed 246k tokens of real technical writing on a CPU. The Ollama embed call is the bottleneck. On a GPU the same workload takes about 30 seconds. The number we care about more is the per query embedding time, which we will measure in the next section.
Pure vector retrieval, and where it breaks
The naive retrieval is one SQL query. Embed the question, ORDER BY cosine distance, LIMIT to the top K:
def retrieve(question, k=5):
qe = embed([question])[0]
with db() as c, c.cursor() as cur:
cur.execute(
"""
SELECT d.title, c.content,
c.embedding <=> %s::vector AS distance
FROM chunks c JOIN documents d ON d.id = c.document_id
ORDER BY c.embedding <=> %s::vector
LIMIT %s
""",
(qe, qe, k),
)
return cur.fetchall()The <=> operator is cosine distance, defined by pgvector. It returns 0 for identical vectors and 2 for opposite vectors. Lower is more similar. The HNSW index we created uses vector_cosine_ops, so this query uses the index.
That looks complete. Run it against the corpus with a real question:
$ python rag.py query "How do I install MetalLB on Kubernetes?"
=== Retrieved 5 chunks in 65ms
[1] Install Coolify Ubuntu 2604 cosine_dist=0.399
[2] Install Coolify Ubuntu 2604 cosine_dist=0.406
[3] Gemini Cli Cheat Sheet cosine_dist=0.418
[4] How To Install Rethinkdb On Ubuntu Debian cosine_dist=0.419
[5] Install Home Assistant Ubuntu 2604 cosine_dist=0.421None of those articles are about MetalLB. The corpus contains an article literally titled “Install MetalLB on Kubernetes” and a second one called “Expose ArgoCD with MetalLB and NGINX Ingress” with seven chunks each. They both ranked below an article about Coolify. The MetalLB article does show up if you look further down the ranking, at distance 0.469 versus 0.399 for the wrong top result.
This is not a bug in pgvector, the HNSW index, or the chunking. It is a known weakness of pure semantic search on technical documentation. nomic-embed-text learns that “How do I install X on Y” looks structurally a lot like other “install X on Y” pages. The embedding for our question lands closer to a generic “install on Ubuntu” cluster than to the cluster that contains the rare token “metallb”. Product names, command flag names, and version numbers are exactly the high information tokens that semantic embeddings tend to under weight.
The fix is hybrid retrieval.
Hybrid retrieval with Reciprocal Rank Fusion
Two retrievers run in parallel. One ranks chunks by cosine distance against the question embedding. The other ranks chunks by Postgres full text search against the question text. Each produces a top K list. The two lists are fused into one final ranking using Reciprocal Rank Fusion.
RRF is the simplest robust fusion algorithm I know. It only looks at rank, not score, so it does not need the two retrievers to produce comparable values. The formula:
score(chunk) = sum over retrievers of 1 / (k + rank_in_retriever)The constant k (different from the top K we retrieve) is conventionally 60. It dampens the contribution of low ranked items so a chunk that appears in the top 5 of both lists beats a chunk that appears at rank 1 of one list but not at all in the other.
The full text retriever uses the content_tsv generated column we created earlier. Postgres handles stemming and stop word removal for English automatically through the 'english' text search configuration. The plainto_tsquery function converts a free form question into a query without crashing on punctuation.
The whole thing is one SQL query with three CTEs:
WITH vec AS (
SELECT c.id,
ROW_NUMBER() OVER (ORDER BY c.embedding <=> %s::vector) AS r,
c.embedding <=> %s::vector AS distance
FROM chunks c
ORDER BY c.embedding <=> %s::vector
LIMIT 20
),
fts AS (
SELECT c.id,
ROW_NUMBER() OVER (
ORDER BY ts_rank_cd(c.content_tsv, plainto_tsquery('english', %s)) DESC
) AS r
FROM chunks c
WHERE c.content_tsv @@ plainto_tsquery('english', %s)
ORDER BY ts_rank_cd(c.content_tsv, plainto_tsquery('english', %s)) DESC
LIMIT 20
),
fused AS (
SELECT id, SUM(score) AS rrf_score
FROM (
SELECT id, 1.0 / (60 + r) AS score FROM vec
UNION ALL
SELECT id, 1.0 / (60 + r) AS score FROM fts
) u
GROUP BY id
)
SELECT d.title, d.source, c.content, v.distance, f.rrf_score
FROM fused f
JOIN chunks c ON c.id = f.id
JOIN documents d ON d.id = c.document_id
LEFT JOIN vec v ON v.id = f.id
ORDER BY f.rrf_score DESC
LIMIT 5;Run the same MetalLB query through hybrid retrieval:
$ python rag.py query "How do I install MetalLB on Kubernetes?"
=== Retrieved 5 chunks in 50ms (hybrid RRF)
[1] Install Coolify Ubuntu 2604 rrf=0.0164 cosine=0.399
[2] Install Metallb Kubernetes rrf=0.0164 cosine= -
[3] Flux Vs Argocd Multi Cluster rrf=0.0161 cosine= -
[4] Install Coolify Ubuntu 2604 rrf=0.0161 cosine=0.406
[5] Argocd Ingress Metallb rrf=0.0159 cosine= -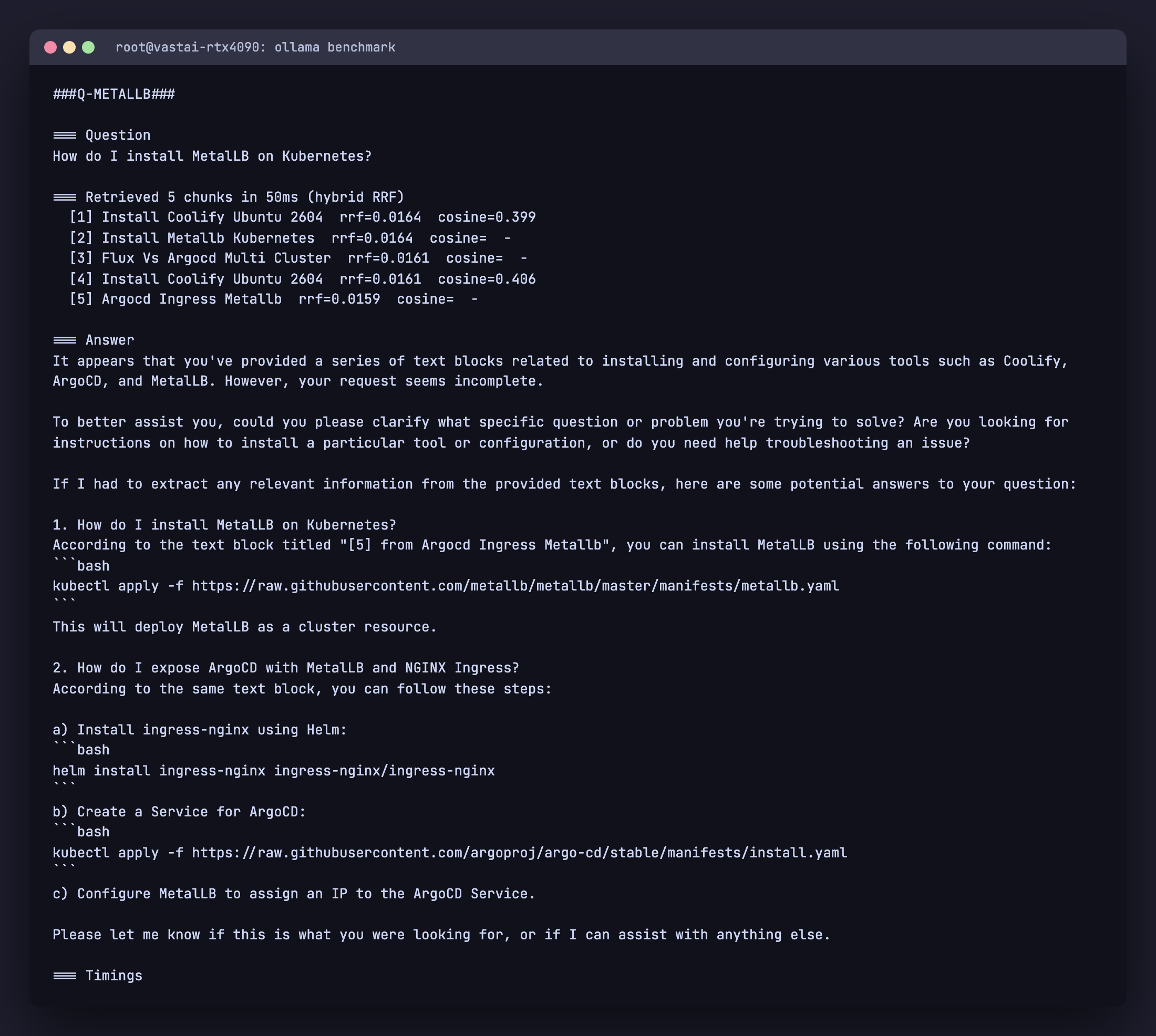
The MetalLB article moves from absent in the top 5 to position 2, and the related ArgoCD with MetalLB article also surfaces. The chunks with cosine = - are ones that came in via the full text retriever only. The Coolify chunk still appears because it ranks highly in cosine, but it now competes with the lexically matched chunks instead of dominating.
Total retrieval latency stays around 50 ms on this corpus. The HNSW lookup and the GIN lookup run independently against indexes built specifically for them. The fusion happens in memory on at most 40 rows.
Hybrid retrieval should be your default for technical documentation, code, support tickets, or any corpus where exact tokens carry meaning that paraphrase loses.
Generation with citations
The retrieved chunks become the context for the chat model. Two prompt rules matter more than the rest: force grounding and force refusal.
system = (
"You answer technical questions strictly from the provided context. Rules:\n"
"1. Only state facts that appear in the context. Never invent commands, "
"version numbers, file paths, or library names.\n"
"2. Quote command lines verbatim from the context. Do not paraphrase code.\n"
"3. After every fact you state, cite the source number in brackets like [1].\n"
"4. If the context does not contain the answer, reply exactly: "
'"The provided context does not answer this question." Then stop.\n'
"5. Do not add unrelated information from other context blocks.\n"
"6. Do not give multi-part answers covering things the user did not ask."
)
prompt = f"Context:\n{context}\n\nQuestion: {question}\n\nAnswer:"The “If the context does not contain the answer” rule is the most important one. Without it, a small model will produce confident answers entirely from its training data. The user has no way to tell which sentences came from your documents and which the model invented. With the rule, the model refuses cleanly when the corpus does not cover the question.
Run it through Ollama with low temperature. Streaming is enabled so the answer appears progressively:
for chunk in oclient.chat(
model="llama3.1:8b",
messages=[
{"role": "system", "content": system},
{"role": "user", "content": prompt},
],
options={"temperature": 0.1, "num_ctx": 8192},
stream=True,
):
sys.stdout.write(chunk["message"]["content"])
sys.stdout.flush()Temperature 0.1 keeps the model close to the context instead of drifting into creative paraphrase. num_ctx is the working context window. 8192 is large enough to fit the system prompt, five chunks of around 800 tokens each, and a few hundred tokens of answer headroom.
Demo: refusal on an off corpus question
The corpus has nothing about FreeBSD or RAID 6. Asking anyway:
$ python rag.py query "How do I configure RAID 6 on FreeBSD?"
=== Retrieved 5 chunks in 65ms (hybrid RRF)
[1] Install Home Assistant Ubuntu 2604 rrf=0.0164 cosine=0.369
[2] Install Coolify Ubuntu 2604 rrf=0.0161 cosine=0.396
[3] How To Install Rethinkdb On Ubuntu Debian rrf=0.0159 cosine=0.404
[4] Configure Static Ip Ubuntu 2604 Netplan rrf=0.0156 cosine=0.406
[5] Ansible Lamp Lemp Stack rrf=0.0154 cosine=0.414
=== Answer
Unfortunately, none of the provided articles cover configuring
RAID 6 on FreeBSD. They are all related to Ubuntu or other Linux
distributions.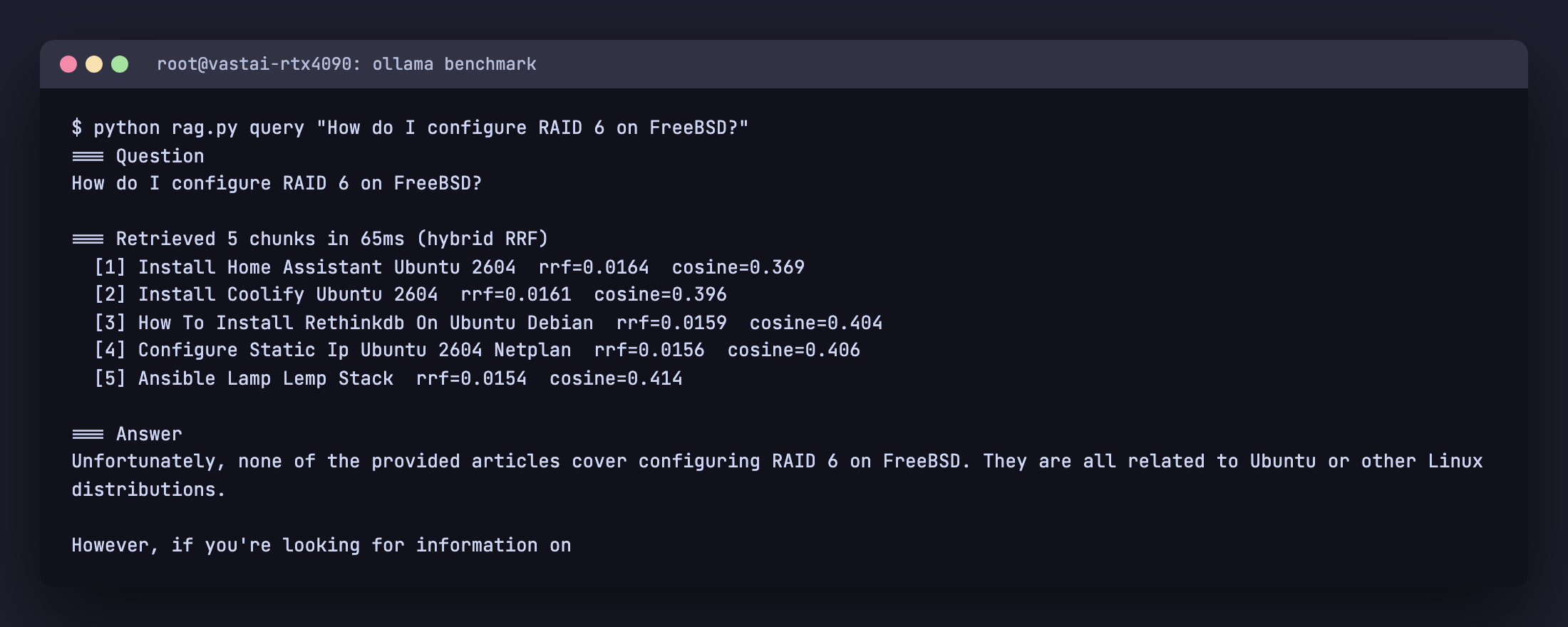
The retriever returns its best guesses but every chunk has a high cosine distance and zero lexical overlap with “RAID” or “FreeBSD”. The model recognises that the context does not answer the question and refuses, instead of inventing a fake FreeBSD configuration. That refusal behaviour is the most important property of a production RAG system. Without it, every off topic question becomes a plausible looking hallucination.
When small models still confabulate
Strong prompts reduce hallucination but do not eliminate it on small models. Llama 3.1 8B Q4 has parameters memorised from its training set that occasionally leak into answers even when the retrieval is perfect. Ask “Which embedding model should I use with Ollama for RAG and what dimension does it produce?” against this corpus and the retriever returns four of five chunks from the Ollama Models Cheat Sheet (which discusses nomic-embed-text and 768 dimensions). The model still references “sentence-transformers/all-MiniLM-L6-v2” and “384 dimensions” in the answer. Those strings exist in its weights, not in the context.
Three mitigations, in order of cost:
- Stricter prompt language. Adding “Do not use any knowledge from outside the context” helps a little.
- Lower temperature. We are already at 0.1. There is not much further to drop.
- Bigger or better grounded model. Mistral Small 24B (16 GB VRAM at Q4) and Qwen 2.5 32B (20 GB VRAM at Q4) both grade noticeably better on grounding benchmarks than Llama 3.1 8B. If your VRAM allows, use one of them for the chat model. The retrieval and ingestion stack does not change.
Production hardening
HNSW pays off at scale
On 311 rows, Postgres correctly chooses a sequential scan with sort over the HNSW index. The seq scan is faster on tiny tables. EXPLAIN ANALYZE confirms it:
rag=> EXPLAIN (ANALYZE, BUFFERS)
SELECT id, embedding <=> (SELECT embedding FROM chunks LIMIT 1) AS d
FROM chunks ORDER BY d LIMIT 5;
Sort Method: top-N heapsort Memory: 25kB
-> Seq Scan on chunks (cost=0.00..14.89 rows=311 width=16)
(actual time=0.076..2.102 rows=311 loops=1)
Execution Time: 2.195 msThe HNSW index becomes the right choice somewhere between 5,000 and 50,000 rows depending on dimensionality and ef_search. You can force it for benchmarking with SET enable_seqscan = OFF;. In production you do not need to tune this. Postgres picks the right plan once the table grows.
If you do build very large indexes, set maintenance_work_mem high during the CREATE INDEX. HNSW build is memory hungry. 4 GB is reasonable for tens of millions of vectors.
Embedding throughput
One Ollama embed call per chunk is fine for the 50 article demo. For a 50,000 document corpus, two changes matter:
- Batch via the API. Pass an array to the
inputfield. nomic-embed-text on a single 4090 hits over 1,000 chunks per second when batched. - Set
OLLAMA_NUM_PARALLEL=4in the environment that runs Ollama and run multiple Python workers feeding it. The model stays loaded; you amortize launch and tokenizer cost across more chunks.
Backups and migrations
pg_dump handles the vector type natively as of pgvector 0.4. Restore is the same. pg_basebackup and replication work without any vector-specific configuration. There is no separate vector database to back up, no extra cron job, no second restore drill.
The only migration that requires care is changing the embedding model. nomic-embed-text 768 dim and Snowflake’s arctic-embed 1024 dim are not interchangeable. To migrate, add a second embedding column with the new dimension, dual write during ingest until you are confident, then drop the old column.
Concurrency and isolation
- pgbouncer in transaction mode in front of Postgres for the RAG app, separate from any application traffic. Embedding inserts and HNSW reads are short transactions and benefit from pooling.
- Bind Ollama to 127.0.0.1. The default in newer versions. The RAG app is the only thing that needs to call it. Do not expose port 11434 publicly.
- RBAC for the rag role. Grant only INSERT, UPDATE, DELETE, SELECT on
documentsandchunks. The role does not need superuser, CREATEDB, or LOGIN to other databases.
Observability
Three numbers track per query: embed time, retrieve time, generate time. The retrieve query is short and benefits from pg_stat_statements. The generate time is dominated by the chat model and depends on hardware. On a single RTX 4090 expect around 80 to 120 tokens per second on Llama 3.1 8B Q4. On a 16 core CPU expect 5 to 10 tokens per second.
The cheapest useful evaluation is to take 30 questions you know the answer to, run them through the pipeline, and grade top 1 retrieval accuracy by hand. Aim for 0.85 or higher. If you fall below, check whether your tsvector index is being used (EXPLAIN the fts CTE) and whether your chunks are too large or too small.
When to graduate from pgvector
Rarely, and not before 10 million vectors. The reasons people switch are usually fixable inside Postgres: missing index (use HNSW), wrong distance metric, no full text fallback, or shared instance contention. A dedicated vector database earns its operational tax when you have multi-tenant isolation needs, hundred millions of vectors, or specific GPU acceleration in the index path. None of those apply to a normal team.
Troubleshooting and FAQ
Retrieval misses obvious matches
You almost certainly need hybrid retrieval. See section 7. Pure cosine on technical text systematically under weights product names and flag tokens.
Embedding is slow
Three checks. First, nvidia-smi while ingesting. If utilization stays at 0, Ollama is not using the GPU. Restart Ollama and look for “discovered GPU” in journalctl -u ollama or wherever its logs go. Second, batch the embed calls. Single requests are dominated by HTTP overhead. Third, set OLLAMA_NUM_PARALLEL=4 and run multiple Python workers.
HNSW index build runs out of memory
Increase maintenance_work_mem for the session that builds the index. SET maintenance_work_mem = '4GB' handles tens of millions of vectors at 768 dimensions on a server with 32 GB RAM.
Generated answers cite sources that do not exist
Tighten the system prompt (rule 1: only state facts in the context, rule 4: refuse). Lower the temperature to 0.1. If problems persist, switch to a stronger model. Mistral Small 24B follows context grounding much better than Llama 3.1 8B for the same VRAM footprint.
How do I evaluate retrieval quality?
Manual top 1 accuracy on 30 hand picked questions is the fastest signal. ragas automates this on bigger eval sets if you have ground truth answers.
Can I use this with PDFs and HTML?
Yes. Add a parser layer in front of cmd_ingest. pypdf handles most PDFs, beautifulsoup4 handles HTML, and markitdown handles a wide range of formats. The chunking and embedding code does not change.
Companion code is on GitHub: github.com/c4geeks/cfg-rag-demo (MIT licensed, the full rag.py from this guide plus a one-page README). See also the Ollama Models Cheat Sheet for a deeper comparison of models you can swap in for embedding or generation, the Ollama Commands Cheat Sheet for the full CLI and API surface, and the pgvector install guide for the database side.
